Amyloids and amyloid-related disorders
Proteins are very important biomolecules that are involved in almost every biological process. In order to function, these biomolecules have to acquire their native structural conformation. Under certain circumstances, however, peptides and proteins can fail to adopt, or remain, in their native functional conformation, and can acquire misfolded conformations that are susceptible to form nonfunctional and potentially harmful aggregates termed amyloids. The onset and progression of more than 50 human disorders including Alzheimer’s disease, Parkinson’s disease, type II diabetes, and prion diseases are associated with the failure of a specific peptide or protein to adopt or remain in its native functional conformational state, and their subsequent conversion into insoluble fibrillar aggregates. In recent years, the process of amyloid formation has emerged as a subject of fundamental importance as it was recognised that many disorders associated with amyloid formation are no longer rare and are rapidly becoming some of the most common medical conditions in the ageing society. Millions of people around the world suffer from amyloid-related disorders, Alzheimer's and Parkinson's diseases alone afflict more than 50 million patients worldwide. Despite significant and sustained efforts, however, the molecular and mechanistic links between protein aggregation and toxicity remain challenging to characterise. In addition, there are still no effective disease modifying drugs or treatment modalities available for amyloid-related disorders. One of the main reasons for this are the complex nature of the peptide and protein aggregation and self-replication, and relatively poor understanding of these processes. Prevention and treatment of a given disease generally require a deep understanding of its underlying causes.
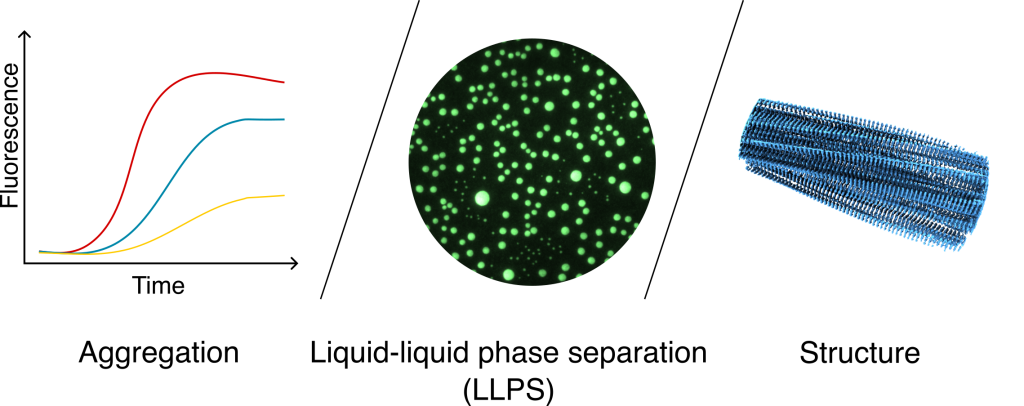
Research areas
Our goal
With our research, we aim to contribute to the greater understanding of amyloid aggregation process. In our research we exploit a variety of biophysical, molecular biology and bioinformatic methods to obtain mechanistic insights into amyloid aggregation process, investigate effects of various environmental factors on the aggregation process and the structure of aggregates. Also, we search for the potential inhibitors of amyloid aggregation.



